Protein backbone angles12/28/2023 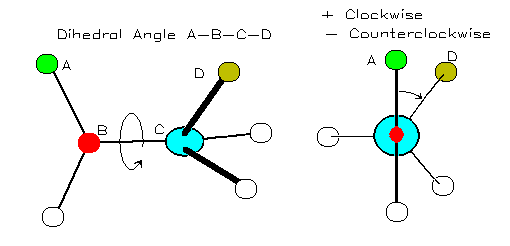 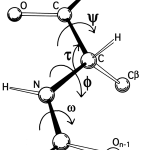
J Magn Reson 202:223233 Clubb RT, Thanabal V. These results highlight the complementarity between AF prediction, solution SAXS and molecular dynamics/conformational sampling for structural modelling of proteins having both structured and flexible regions. backbone amide resonances with the alpha-carbon of the preceding residue in. A fast ensemble modelling method was employed that optimizes the fit of pair distance distribution functions and intensity profiles computed from the pool to their experimental counterparts. A pool of representative structures was generated using a Monte Carlo method that adjusts backbone dihedral allowed angles along potentially flexible regions. Wu et al.proposed a composite machine-learning algorithm called ANGLOR to predict real-value protein backbone torsion angles from protein sequences. Instead, excellent agreement is found with ensemble models generated by allowing for flexible linkers between high-confidence predicted structured domains. The method is expected to be useful for advancing protein structure and function prediction. A number of methods, mostly data-driven machine learning methods, have been developed to predict torsion angles from protein sequences and the predictions can then be used as restraints to predict. In particular, 44 of 40-residue fragment structures constructed from predicted backbone C-based and angles are less than 6 Å root-mean-squared-distance from their native conformations, nearly 20 better than the next best. Three examples were identified where there is clear evidence that the single AF-predicted structure cannot account for the experimental SAXS data. The conformation of the backbone of a protein can be largely represented by two torsion angles (phi and psi angles) associated with each C atom. The backbone structure of a protein can be determined by its torsion angles (see Figure 1 for illustration of what Psi Phi angles are). Selected SASBDB entries were carefully examined to ensure that they represented data from monodisperse protein solutions and had sufficient statistical precision and q resolution for reliable structural evaluation. Focusing on single-chain structures without prosthetic groups, an earlier comparison of features derived from small-angle X-ray scattering (SAXS) data taken from the Small-Angle Scattering Biological Data Bank (SASBDB) is extended to those calculated using the corresponding AF-predicted structures. While a stunning accuracy has been achieved for many folding units, predicted unstructured regions and the arrangement of potentially flexible linkers connecting structured domains present challenges. By providing predicted protein structures from nearly all known protein sequences, the artificial intelligence program AlphaFold (AF) is having a major impact on structural biology.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
